Welcome
News
Our assembly & annotation software LRSDAY got published!
The long-read-based yeast genome assembly and annotation software LRSDAY is officially out in Nature Protocols!
You can read the paper via this link: https://www.nature.com/articles/nprot.2018.025
The software is hosted at GitHub: https://github.com/yjx1217/LRSDAY

Nice Matin reported our work!
Our work is highlighted by the regional newspaper Nice Matin today!

CNRS INSB hilighted our work!
Our work is highlighted by CNRS - Institut des sciences biologiques (INSB) today: http://www.cnrs.fr/insb/recherche/parutions/articles2017/g-liti.html
Paper Published!
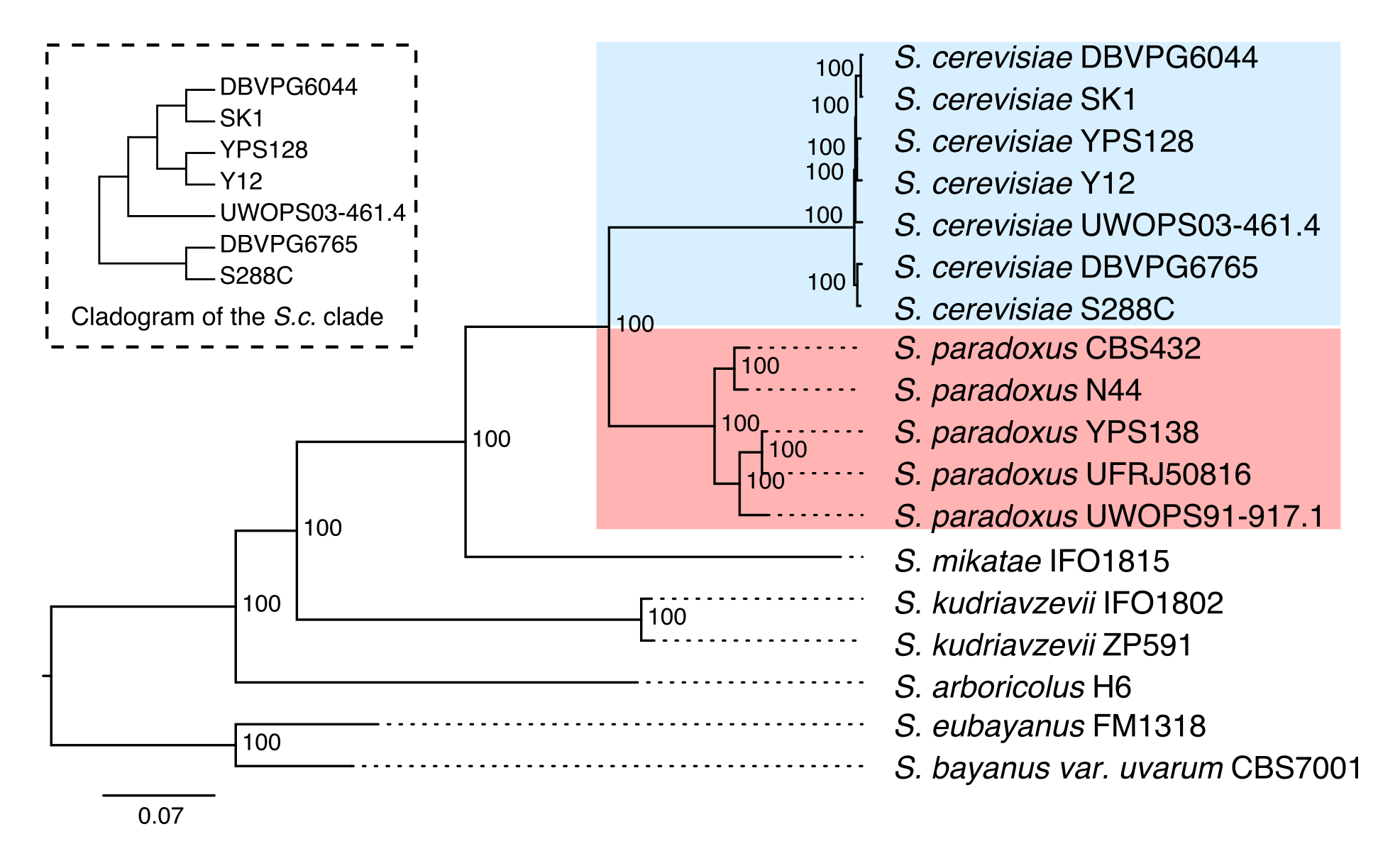
Our paper “Contrasting evolutionary genome dynamics between domesticated and wild yeasts” is officially out in Nature Genetics with open access! You can read the paper via the following link: http://www.nature.com/ng/journal/v49/n6/full/ng.3847.html

Data update
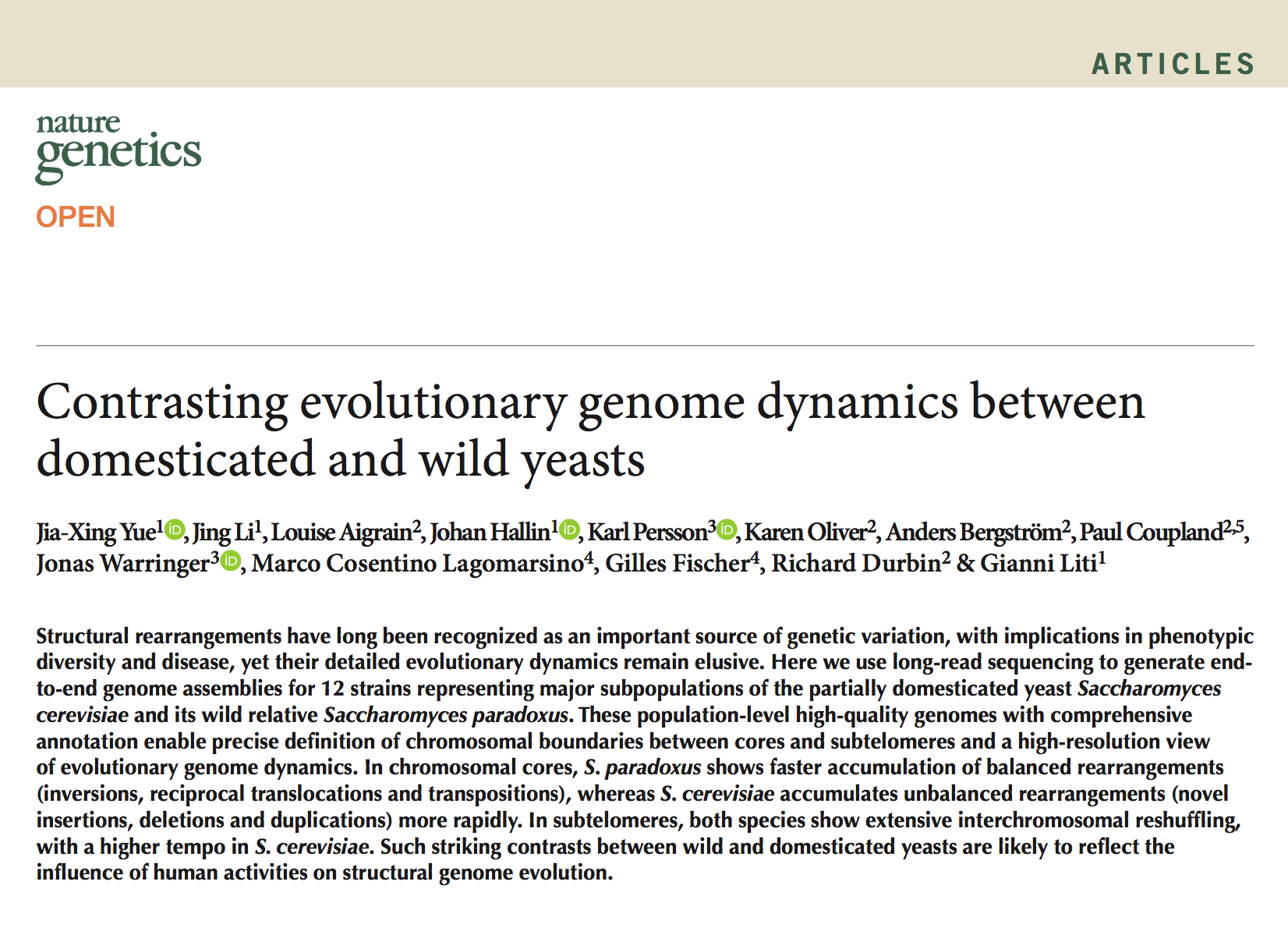
Please note that the mitochondrial genome assemblies and annotations of S. paradoxus UFRJ50816 and UWOPS91-917.1 were updated on Thu Mar 16 06:17:38 CET 2017. If you have downloaded the corresponding data before this update, please use the new data for your analysis.
Data update
Please note that we made minor improvements for both the assembly and annotation data today. If you have downloaded the corresponding data before this update, please use the new data for your analysis.
Welcome!
We set up this website to provide general description and data release for the Yeast PacBio Sequencing project. In this project, we sequenced 12 strains representing major subpopulations of two close-related Saccharomyces yeast species: S. cerevisiae and S. paradoxus. We feel the high-quality genome assemblies and annotations that we generated in this study will be of great interests to the yeast community. We want to share this valuable dataset to the community to facilitate future genomic and functional investigations on this important model organisms. The preprint of this work can be found here.
subscribe via RSS